INDIVIDUAL RESEARCH PROJECT 2
OMERO embedding and Optimisation of Biom3d, an image analysis tool using deep-learning networks
- Home /
- Individual Research Projects /
- OMERO embedding and Optimisation of Biom3d, an image analysis tool using deep-learning networks
Abstract
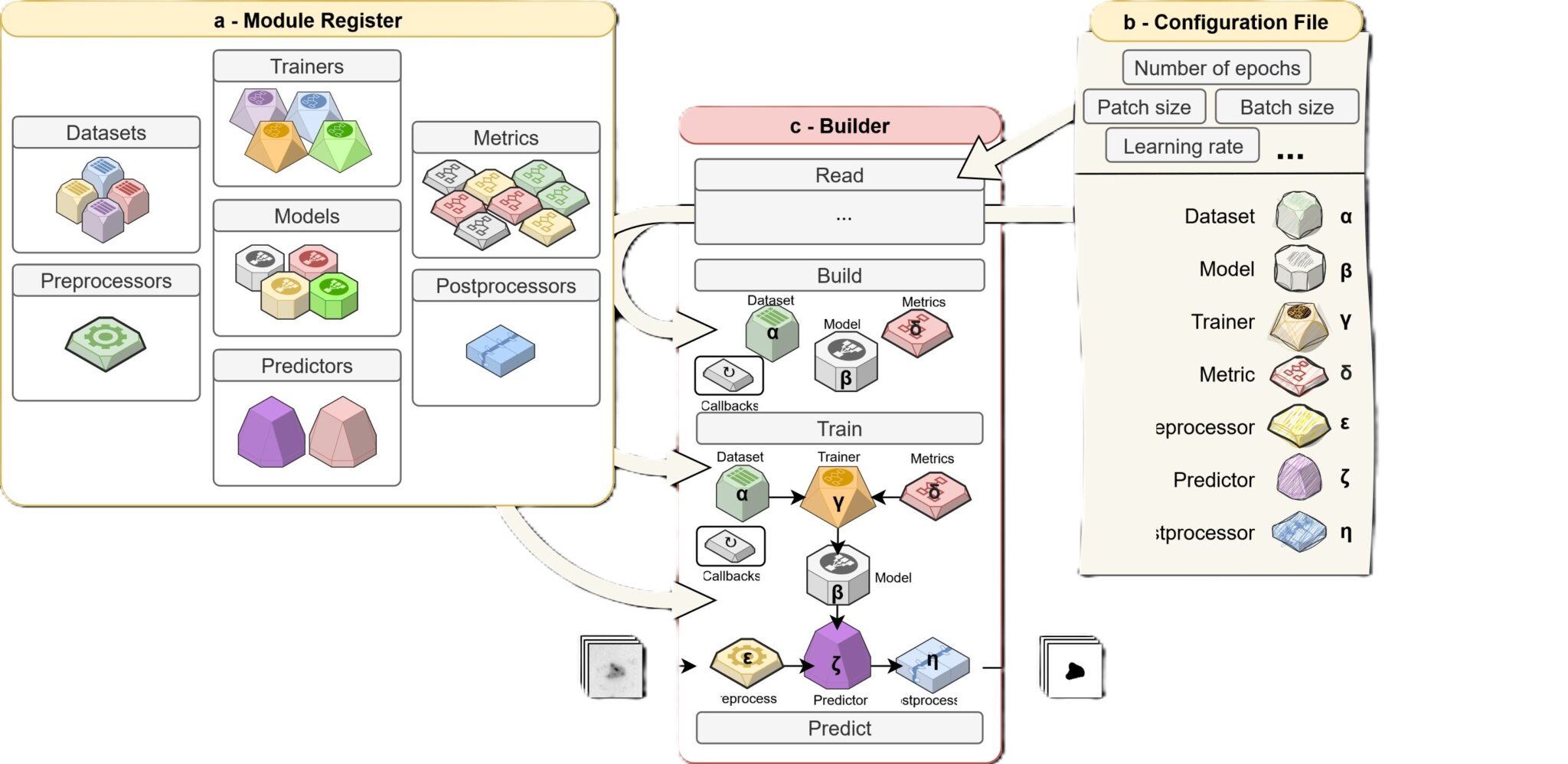
Neural networks, employing deep learning (DL), have become essential for high-throughput image analysis. However, DL tools should be made accessible to a broader scientific community, including plant biologists. In line with the FAIR (Findable, Accessible, Interoperable, Reusable) principles, (Obj1) we aim to implement new image analysis functionalities into Biom3d, a modular framework based on DL networks initially designed to quantify nuclear shape and chromatin organization.
(Obj2) To facilitate the use of Biom3D for end users’, we will improve its interoperability with OMERO, a popular image data management system created by the University of Dundee, with Imaris, a proprietary software from Oxford Scientific, and with the FRACTAL web-platform developed by the bioimaging center in Zürich in order (Obj3) to contribute to the design of a new sets of workflows called ChromTrack3D adapted to microscopists’ needs in image analysis. Finally, we will explore connecting OMERO servers between the AGILE consortium partners to promote the accessibility and sharing of image datasets in collaboration with the University of Dundee.
More information
Training benefits
Developing a new generation of modulable frameworks to perform quantitative image analysis is central for microscopists. These frameworks like Biom3d and FRACTAL used in this project will be adapted to use images coming from various species, modalities (eletron, confocal or light sheet microscopy), or image formats (like OME-Zarr) and thereby made interoperable with data management systems like OMERO for best reproducibility.
You will learn how to implement and train new deep-learning models in the Biom3d framework, develop OMERO interface scripts, run jobs on High Performance Computing (HPC) clusters and develop new functionalities in Biom3d like preprocessing OME-Zarr images.
Requirements
Significant experience in Python (and Java) scripting, and knowledge in (plant) molecular biology. A Master in the bio-informatics or image analysis. A desire to succeed in science.
Environment
Clermont-Ferrand (France) is a middle size city with a high quality of life, a stimulating scientific environment, and the city is bordered by the Chaine des Puys, a UNESCO classified volcanic formation with great outdoors activities. The Chromatin and 3D Nuclear Dynamics (CODED) team co-lead by Aline Probst and Christophe Tatout is a pluri-disciplinary team working in plant epigenetics and bioimage analysis in a friendly working environment. The team has an international recognition and is leading the network EPIPLANT of the French epigenetic community.
Responsible PI


