INDIVIDUAL RESEARCH PROJECT 14
Characterisation of peripheral light-induced transcription clusters
- Home /
- Individual Research Projects /
- Characterisation of peripheral light-induced transcription clusters
Abstract
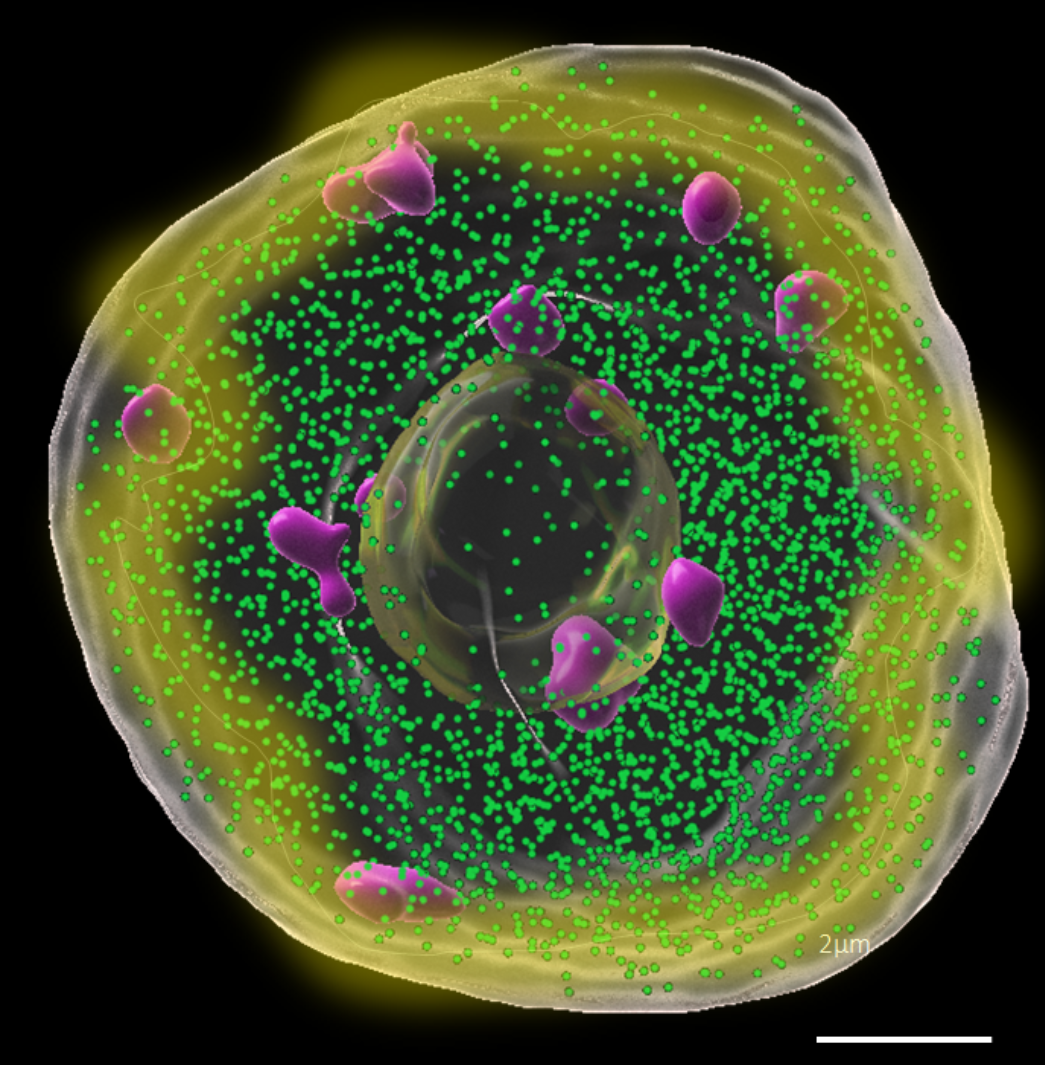
The three dimensional arrangement of the genome in the nuclear space likely plays a significant role in transcriptional regulation. Our goal is to understand whether and how the 3D organisation of transcription contributes fast responses to the environment. Using 3D STED imaging we identified Light Induced Transcription Clusters (LITC) enriched at the nuclear periphery of plant cells exposed to light. The project aims at: (Obj1) quantifying LITC distribution relative to the nuclear pore complex, in wild-type and in mutants lacking a functional plant lamina, (Obj2) determining LITC proximal chromatin environment relative to nucleosomes (H2B), linker histones (H1) and epigenetic domains (H3K4me3, H3K27me3).
To address these questions the candidate will deploy in-house protocols for intact nuclei preparation, immunostaining, 3D STED imaging and image analysis but will also work on improvements by testing new STED-optimized fluorescent dyes in collaboration with Abberior (secondment1) and by creating re-usable Imaris Workflows in collaboration with Bitplane (secondment 2). Finally, with the long term aim to obtain a 3D map of gene expression in the nucleus, the candidate will work towards establishing a multiplex in situ FISH (MERFISH) approach (Obj3) in collaboration with the Earlham Institute (secondment 3).
More information
Training benefits
Learning and applying cell biology (nuclei isolation, embedding, immunostaining, FISH), bioimaging at super resolution imaging (STED) and image analysis including segmentation and image data analysis. The student will enroll in one of the Plant Science Center or Molecular Life Science doctoral programmes (www.lifescience-graduateschool.uzh.ch) offering modules complementary to AGILE’s training programme.
Requirements
Experience with fluorescence microscopy, molecular and cell biology. Strong interest in combining experimental and computational work, not afraid of repetitive tasks and troubleshooting. Curious mind and interest in problem solving. Master degree in Cell and/or Molecular Biology, or Master in Bioimaging.with interest in learning basic molecular biology.
Environment
The University of Zürich (https://www.uzh.ch/) offers a vibrant and international environment for students with excellence level of doctoral training, and high-end infrastructure for research, including in bioimaging (zmb.uzh.ch ) and advanced image analysis (biovisioncenter.uzh.ch). Doctoral students at our department (Plant and Microbial Biology) have a highly active community offering support and organising various leisure activities beside work. Zürich itself is a “little city with big charm”, with a high quality of life, close to nature, and well connected to other close-by cities (Basel, Bern, Geneva).


