INDIVIDUAL RESEARCH PROJECT 7
Characterizing altered 3D chromatin organization in Arabidopsis pds5 mutants
- Home /
- Individual Research Projects /
- Characterizing altered 3D chromatin organization in Arabidopsis pds5 mutants
Abstract
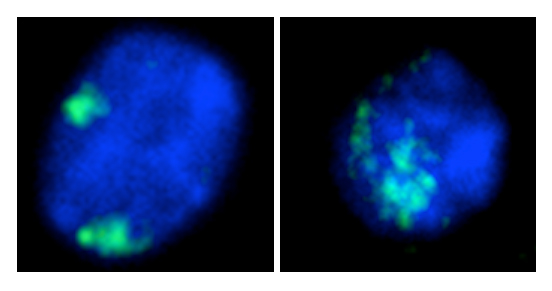
This project focuses on characterizing changes in spatial chromatin organization in the Arabidopsis pds5 mutants that alter chromatin compartmentalization and form prominent Topologically Associating Domains (TADs) across the genome. Using ploidy-sorted nuclei, which allows us to exclude potential variations amongst nuclei due to endoreplication, we aim to achieve the following objectives: (Obj1) with FISH, to measure the distance between lamin-associated domains (LADs) and the nuclear periphery in pds5; (Obj2) with immunostaining, to characterize the spatial distribution of nuclear lamin proteins in pds5; (Obj3) with SABER-FISH, to measure the spatial proximity between genomic regions located within newly emerged TADs in pds5 nuclei, and compare it to wild-type. (Obj4) With the ANCHOR system, to quantify the mobility of candidate chromatin loci in pds5.
More information
Training benefits
Skills in advanced confocal imaging with Airyscan 2
Acquisition of State-of-the-art techniques about three-dimensional plant chromatin organization
Secondment opportunities to learn live chromatin imaging and spatial transcriptomics techniques
Requirements
Good background knowledge of functional genomics and epigenetics
Lab experience in molecular experiments concerning chromatin
You have your own ideas after reading our recent paper DOI: 10.1038/s41467-024-53760-x
Environment
Stuttgart is a global center for innovation with rich culture, green spaces, and industry connections. It provides an inspiring academic environment and outstanding opportunities for personal and professional growth. The University of Hohenheim, located in this great city, offers cutting-edge research, international collaboration, and excellent support for PhD students.


