INDIVIDUAL RESEARCH PROJECT 1
Chromatin mobility induced by temperature changes
- Home /
- Individual Research Projects /
- Chromatin mobility induced by temperature changes
Objectives
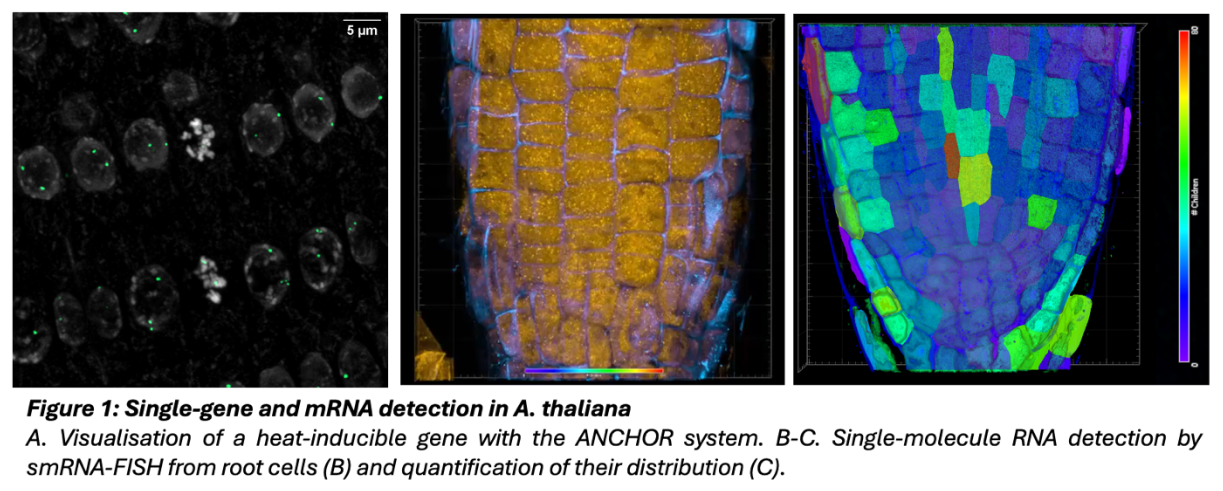
Heat stress affects heterochromatin organization and nuclear movement, but the impact remains unknown at the gene scale. This IRP investigates how nuclear and chromosomal positioning affect gene mobility and transcriptional activity. Measure will be performed under control and heat stress conditions, in wild-type and in plants impaired in two mutants affected in temperature sensing and adaptation (hsfA2 and hsp101). This project will greatly benefit from a comprehensive collection of Arabidopsis thaliana lines tagged at hundreds of individual locations in the genome using the ANCHOR single-copy monitoring system created by our team.

(Obj1) Quantify the mobility of 12 chromatin regions labelled with the ANCHOR system during heat wave response (27°C, 37°C) using ChromTrack3D.
(Obj2) Quantify the chromatin mobility of three candidate loci responding to heat stress (27°C, 37°C) in wild-type and in plants affected by the heat stress response using ANCHOR and ChromTrack3D.
(Obj3) Quantify the expression changes of the three candidate loci with an allelic resolution using the MS2 tag and single molecule RNA FISH coupled with ANCHOR.
In parallel, a collaboration with informatician and the NEoVirTech company (S1) will be realized to optimize our existing gene tracking method for long-term (up to 12 h) live imaging enabling chromatin mobility measurements in 3D (ChromTrack3D). In addition, the use of microfluidic devices will be implemented to reduce the stress on plants generated by long-term experiments (S2).
More information
Training benefits
- Conceptual and practical training in epigenetics/chromatin regulation.
- Hands-on training in advanced confocal microscopy, live-cell imaging, and single-molecule RNA imaging.
- Image analysis using modern AI-based tools for segmentation, tracking, and quantification.
- Structured teaching and mentoring experience with feedback on pedagogy and communication.
Requirements
Applicants must hold a Master’s degree, have basic molecular biology skills, plant genetics and a strong interest in cell biology/microscopy. Previous experience in confocal microscopy would be a plus.
Environment
Actually in Perpignan (LGDP, France), Frédéric Pontvianne’s team will join Clermont-Ferrand (France) and the Chromatin and 3D Nuclear Dynamics (CODED) team co-lead by Aline Probst and Christophe Tatout is a pluri-disciplinary team working in plant epigenetics and bioimage analysis in a friendly working environment. The team has an international recognition and is leading the French epigenetic community and is working at the iGRED institute, a stimulating scientific environment. Clermont-Ferrand (France) is a middle size city with a high quality of life, surrounded by magnificent volcanoes.
Responsible PI

Coordinator of the AGILE consortium
ORCID link :
0000-0002-2913-4104
Website links :
Actual Website
Future Team Website

